IV) Computer Assisted Drug Design
A) Computer Assisted Drug Design
1. Design of Non-toxic Enediyne Antitumor Drugs
Microorganisms and human beings have one thing in common: Both are attacked by toxic bacteria and viruses. However, microorganisms are on earth at least two billions of years longer than human beings and, therefore, they know much better how to protect themselves against bacteria and viruses. Naturally occurring enediynes (calicheamicin, esperamicin, etc.) produced by microorganisms have the ability of destroying the DNA of toxic bacteria and viruses in a three-step process. First, a delivery system helps the enediyne to dock into the minor grove of the DNA helix. Then, a molecular trigger device causes the enediyne unit of the natural compound to undergo a Bergman cyclization and to form a highly reactive p-didehydrobenzene biradical, which is positioned in the minor grove in such a way that it can abstract two proximal hydrogen atoms from the deoxyribose sugars on both strands of the DNA, thus damaging DNA to form a stable arene. This leads to cleavage of the DNA at both strands and, accordingly, causes cell death.
This simple three-step mechanism, which is highly effective in nature, has motivated many scientists to investigate whether such a mechanism could be used to destroy the DNA of tumor cells. However, the problem with the naturally occurring enediynes is that they cannot differentiate between tumor and normal cells in the human body, i.e. all naturally occurring enediynes are highly toxic. - In our previous work on enediynes anticancer drugs, we concentrated on the correct description of the Bergman reaction and the characterization of the organic biradicals formed, which are the key compounds for understanding the unusually high biological activity of enediynes. Some of the highlights of our previous work can be summarized as followed.
- 1995First reliable description of the Bergman reaction by CCSD(T) calculations results of which were later fully confirmed by kinetic measurements of Roth and co-workers.
- 1996First identification and characterization of the m-didehydrobenzene biradical by CCSD(T) calculations and low temperature matrix isolation spectroscopy. This work was highlighted in Chemical Engineering News, 8, 36 (1996).
- 1998First identification and characterization of the p-didehydrobenzene biradical by DFT calculations and low temperature matrix isolation spectroscopy.
- 1999Proof and explanation why DFT with the B3LYP approach provides a reasonably accurate description of the Bergman reaction and didehydroarene biradicals.
- 2000Setting up the requirements, which a new enediyne antitumor drug must fullfill.
- 2000Suggestion of a new enediyne warhead.
In 2000, we developed seven criteria for the design of a new enediyne compound that meets the requirements of an efficient antitumor drug being highly active, selective, and non-toxic. The latter criterion will be fulfilled using the fact that the medium of the tumor cell is slightly acidic (M. von Ardenne, Adv. Pharmacol. Chemot. 109, 339, 1972; see, also M. Vol'pin, patent A61K 31/70, 31/245, 33/24 ) where the acidity can be increased by invoking hyperglycemia. According to our strategy, the ideal enediyne warhead should contain a heteroatom such as N, which can be protonated in the acidic medium of the tumor cell (this implies a large pKa value of the hetero-enediyne). Whereas in the neutral medium of the normal cell the hetero-enediyne is hardly active (Bergman rearrangement is slow, the kinetic stability of the intermediate biradical is low so that it cannot develop its H abstraction ability) the protonated form in the tumor cell shows increased biological activity (the Bergman reaction becomes fast; the biradical character and by this the H abstraction ability increase; the kinetic stability of the biradical is high).
Utilizing and improving the criteria worked out so far, we carried out a systematic description of hetero-enediynes. The project comprised the following steps: 1) Calculation of reaction profiles for Bergman and retro-Bergman reactions of a series of hetero-enediynes using DFT, CCSD(T) and BD(T). – 2) Calculation of singlet-triplet splittings, determination of biradical character; calculation of H abstraction reactions using suitable DNA models such as THF, furanose, etc. – 3) Determination of the pKa values for the molecules being active in the cell. - 4) Large scale DFT calculations of the total drug with the new endiyne warhead found in 1-3.
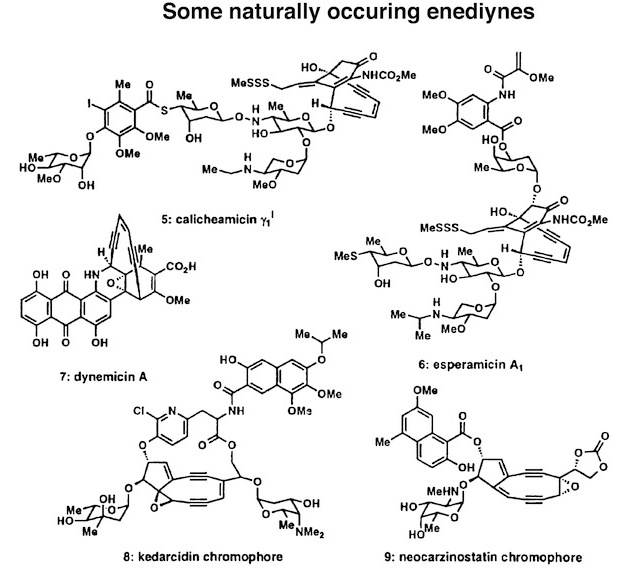
Figure 1. Some naturally occurring enediynes and their discovery details.
2. Interaction of Dynemicin with DNA
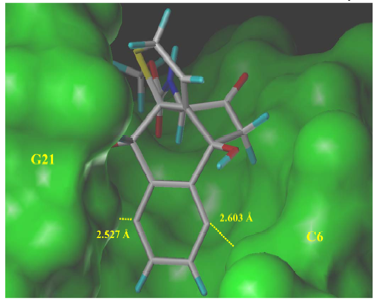
We studied the naturally occurring enediyne Dynemicin A by means of quantum chemcial calculations and molecular modeling. We could determine structure, conformation, and biological activity of dynemicin A. In particular, we were able to describe for the first time geometrical and energetical details of the docking and triggering mechanism of this compound as it happens in the minor groove of DNA. We could clarify that the H abstraction proceeds via a two step mechanism.
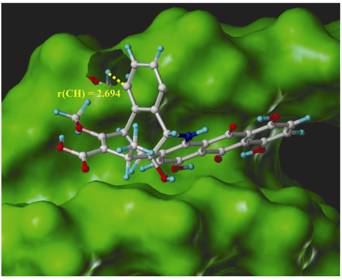
Figure 2: Hydrogen abstraction G7-H5' by Dynemicin A modeled in the minor groove of the duplex 10-mer B-DNA sequence d(CTACTACTGG)-d(CCAGTAGTAG)
Based on this detailed knowledge, a modified dynemicine A was designed, which fulfills the criteria for non-toxicity. The drug obtained in this way does not attack DNA in the normal cell because its warhead is destroyed at pH 7. In the tumor cell, it is activated at body temperature and attacks the DNA in the cell.
The research carried out focused on dynemicin because it is the simplest of all naturally occurring enediynes. The following steps were performed:
- We investigated the Bergman cyclization of the dynemicin at the DFT level of theory to evaluate how mechanism and especially the reaction barrier compare with the parent enediyne and 10-membered ring model systems. The singlet-triplet splitting (which determines the biological activity of an enediyne) of the dynemicin biradical was evaluated and compared with that of suitable model systems.
- We used the known X-ray structure of dynemicin to investigate how its geometry differs in the solid state, in the gas phase, and in aequous solution where in the latter case we applied a continuum model based on DFT.
- Especially, we investigated the triggering mechanism of dynemicin, which leads to the Bergman reaction and the formation of the biradical. We determined under which physiological conditions the Bergman cyclization of dynemicin is triggered.
- Following this work we designed a new dynemicin drug by merging the naturally occurring enediyne with the amidine warhead investigated in the previous project. The resulting compound should be a suitable candidate for a non-toxic antitumor drug provided the properties calculated for the amidine do not change strongly under the impact of trigger device and delivery system.
- Finally, we investigated the in vivo situation by modeling the dynemicin in the minor grove of DNA with the help of QM/MM where the QM part is based on DFT and the MM part on MM2 calculations.